The Publication Workbench is a separate window in Sensitify Studio for assembling publication-grade figures across experiments: calibration curves, Randles–Ševčík analyses, Tafel slopes, dose-response sigmoids, and distribution comparisons. The main Studio window handles one experiment at a time. Figures export to PDF, PNG, or SVG.
Sensitify built it to replace the round-trip through Origin, Excel, or Matlab that most labs run after every measurement campaign.
Open the Workbench
The main Studio toolbar has a Publication button (chart icon) that opens the Workbench in a new window. Studio stays running behind it.
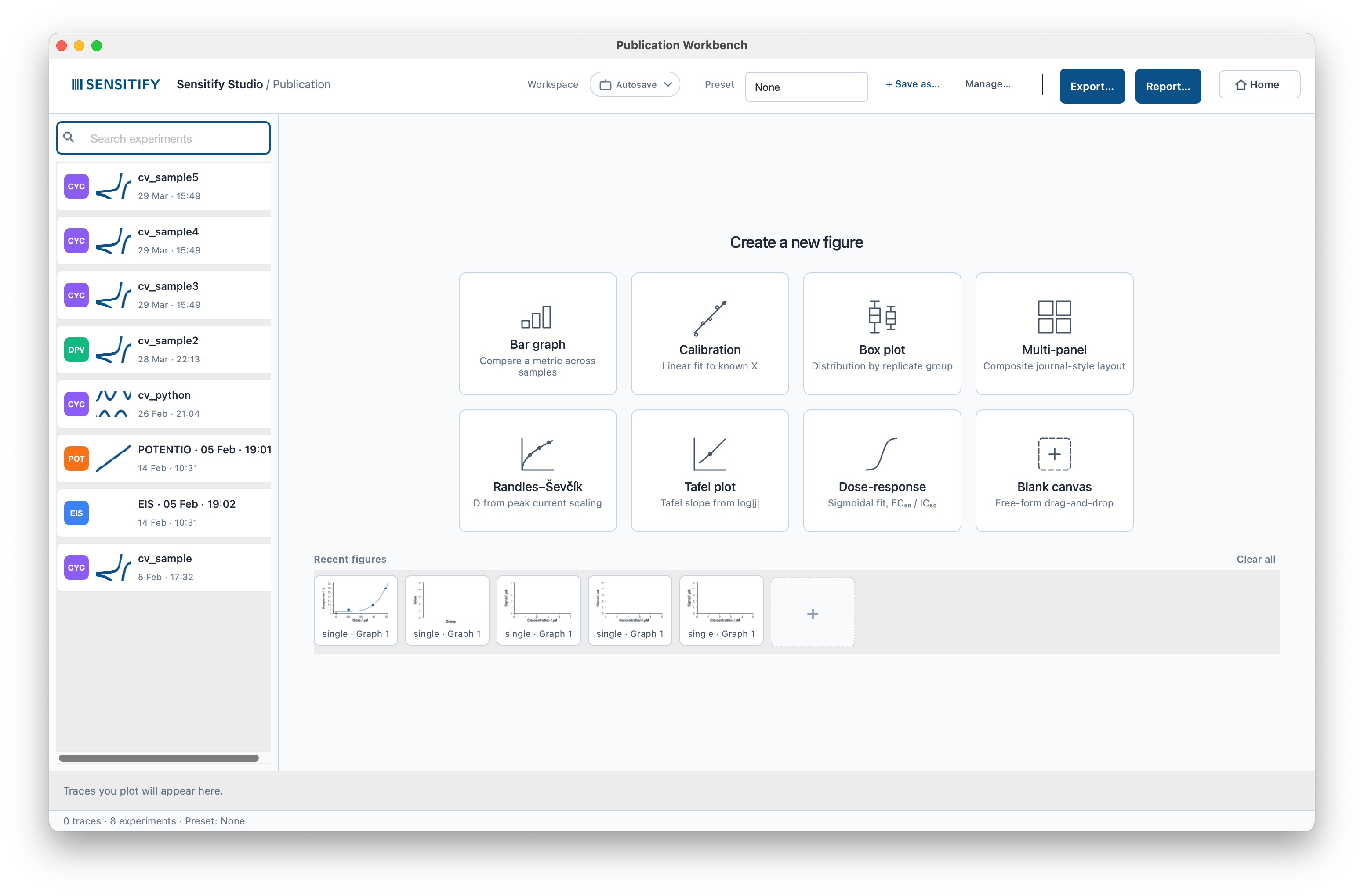
The window at a glance
- Top toolbar. Workspace and Preset selectors, Save as… / Manage… for the active preset, Export, Report, and a Home button back to the main Studio window.
- Left rail. Every experiment in the current workspace, tagged by technique. Cmd/Ctrl-click to multi-select; drag experiments onto a figure or right-click to start an analysis from the selection.
- Centre. The welcome card grid when no figure is open, the figure canvas while you work, and a Recent figures strip at the bottom for quick re-entry.
- Right inspector. Figure-grade style controls: scatter colour and size, fit-line width, CI band opacity, marker visibility, font and size, log-axis toggles.
- Bottom status line. Trace and experiment counts and the active preset name.
You can launch an in-canvas analysis from three places: the
header + Calibrate / + Tafel buttons, the welcome card
grid, or the library right-click menu.
In-canvas analysis modes
Instead of a modal dialog, an in-canvas analysis mode takes over the left rail with a data editor while the canvas renders the live figure. Edit a value, and the fit re-runs within 100 ms.
Each mode shares the same components:
- A 5-column data table (include, source/label, X, Y,
delete) with
From experiments/Manual entrytoggle, paste from clipboard, and CSV / TSV import. - A live fit / stats panel that recomputes after every edit.
- Inline validation. Yellow banners warn but allow; red blocks Done.
- Cmd+Z undo, 32 deep, scoped to the mode.
- Named presets per mode (
Save as…/Manage…) so a setup like "MIP-glucose Mar 2026" is one click to reload. - An Inspector panel on the right for figure styling.
Calibration
Linear regression of concentration (X) against measured signal
(Y). Reports slope, intercept, R², RMSE, LOD (3.3 σ_blank /
|slope|), and LOQ (10 σ_blank / |slope|). Type a measured
signal into the Predict: field and the Workbench
back-calculates the concentration with a 95 % prediction
interval. This single feature replaces a multi-step Excel
exercise. A concentration unit picker (nM / µM / mM / M / mg·L⁻¹ / ppm / ppb) drives the axis label, slope unit, LOD
unit, and inverse-predict result unit together.
Replicate rows (same concentration, different Y) collapse to
one mean point with an SD error bar on the plot, but every
replicate is used in the fit so the degrees of freedom stay
correct. Rows labelled blank (case-insensitive) feed σ_blank
automatically; the intercept SE is the conventional fallback.
Outliers (residual > 2 σ) get an orange marker, but the
Workbench never auto-excludes them. The user decides.
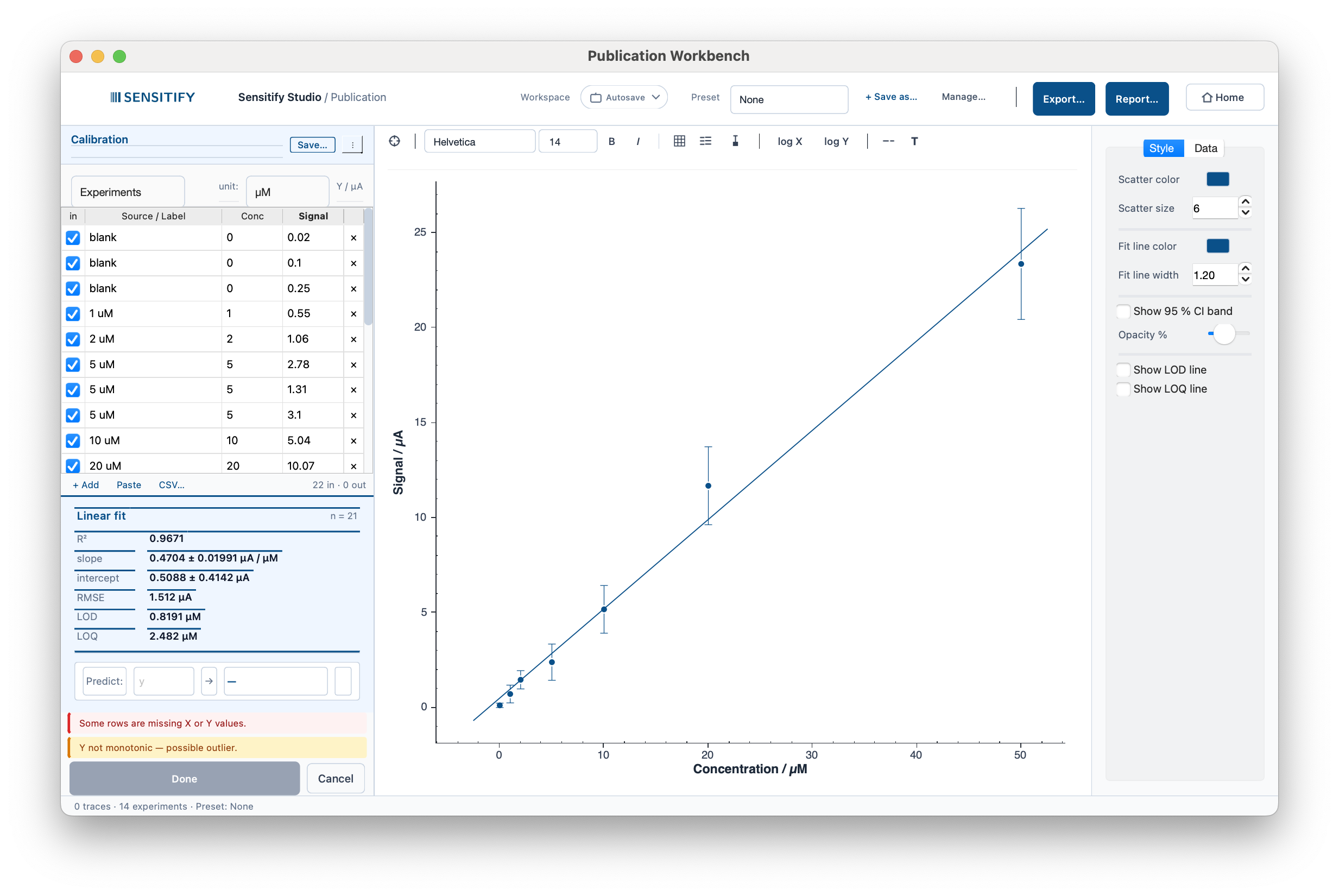
A dopamine MIP biosensor calibration by DPV, 0–50 µM linear
range with three blanks and a triplicate at 5 µM. Slope ≈
0.50 µA/µM, R² ≈ 0.9999, LOD ≈ 0.16 µM. The triplicate
collapses to one mean point with an SD error bar, and σ_blank
is read automatically from the three rows labelled blank.
Randles–Ševčík
Cyclic-voltammetry scan-rate analysis. The Workbench plots
|i_p| against √ν, fits a line, and solves the Randles–Ševčík
equation for the diffusion coefficient D in cm²/s with proper
unit conversion (X in V/s or mV/s, Y in nA through A). Standard
error is propagated end-to-end.
Spinboxes for n (electrons), A (electrode area), C (bulk concentration), and T (temperature) live in the editor. Change any of them and the fit re-runs instantly.
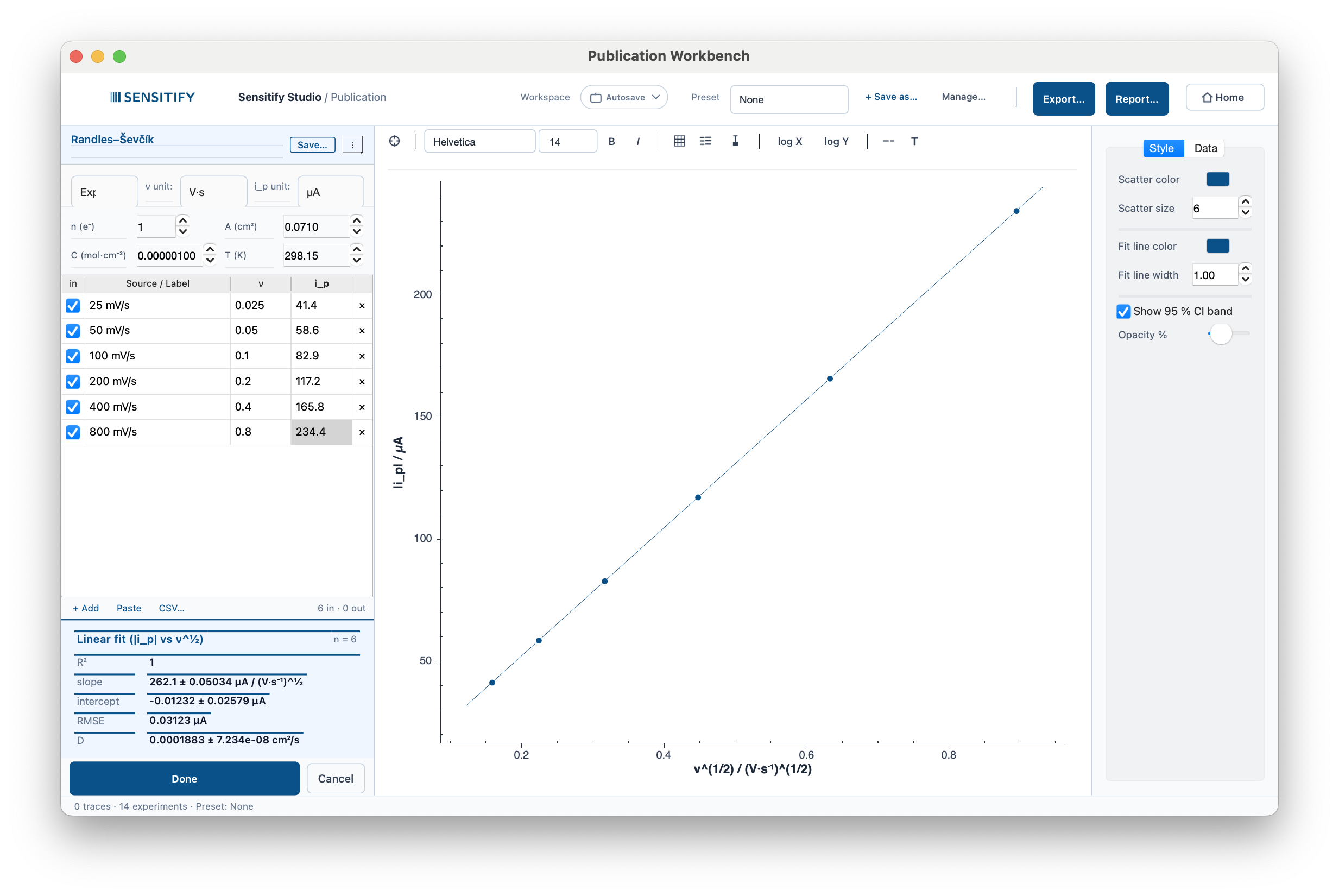
5 mM K₃[Fe(CN)₆] / 0.1 M KCl on a 3 mm GC electrode, six scan rates from 25 to 800 mV/s. The Workbench plots in (√ν, |i_p|) space and reports the diffusion coefficient natively: D ≈ 7.6 × 10⁻⁶ cm²/s, R² ≈ 0.9999. Standard error is propagated end-to-end through the unit conversion.
Tafel
log|j| vs E plot with separate linear fits per branch. The slope of each branch is the Tafel slope β in mV / decade regardless of whether the user typed potential in V or mV; the intercept gives the exchange current density j₀.
The fit-window edges render as dotted vertical lines on the plot, green for the anodic branch and red for the cathodic, so you can see which points each fit uses. Drag the spinbox edges to widen or narrow either window and watch β update live. Disable the cathodic branch with one toggle when the negative side is noisy. Rows where |j| = 0 are skipped automatically (log undefined) with a yellow warning naming the rows.
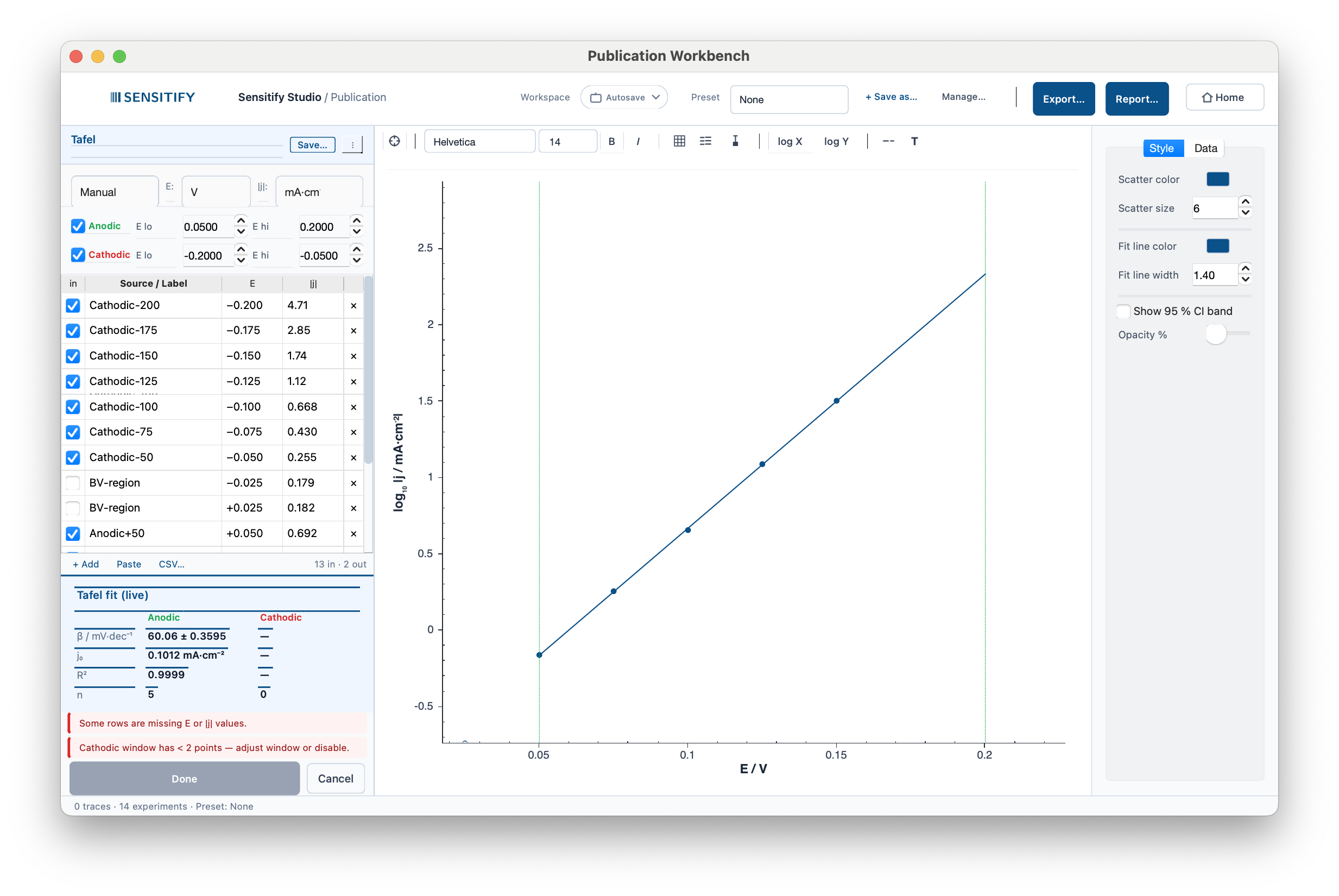
A mild-steel coupon in 1 M HCl, LSV around the corrosion potential. The two branches fit live: β_a ≈ 60 mV/dec (anodic dissolution), β_c ≈ 120 mV/dec (hydrogen evolution), j₀ ≈ 0.10 mA/cm² where the extrapolations meet. The green and red dotted lines are the anodic and cathodic fit-window edges. Drag a spinbox to refit in real time.
Dose-response (4-PL)
Four-parameter logistic fit: y = bottom + (top − bottom) / (1 + (x / EC₅₀)^hill). Extracts EC₅₀ / IC₅₀, the dose at half-max response. The solver is Levenberg–Marquardt with an analytical Jacobian and quintile-based initial guesses; convergence is robust across typical pharma data shapes.
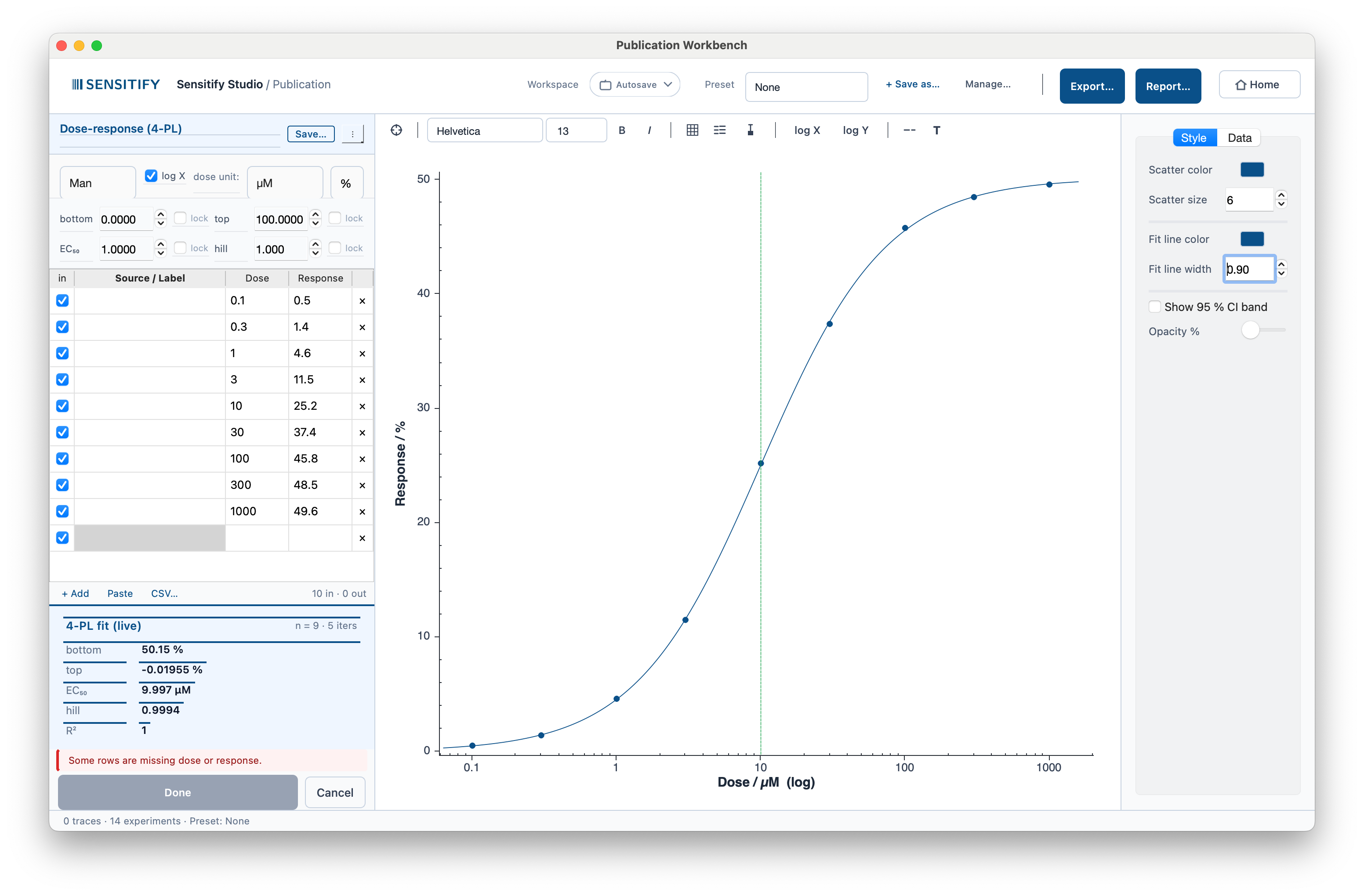
A 4-PL fit through nine dose / response points spanning five decades. The Workbench recovered EC₅₀ ≈ 10 µM, Hill slope = 1, top ≈ 50 (units in the response axis), R² = 0.9994. The vertical line marks EC₅₀.
Per-parameter locks let you fix bottom at 0 (when
background is known to be zero) or top at 100 % (full
saturation) and refit only the remaining parameters. Use this
when you need tighter EC₅₀ confidence intervals on
saturating-response assays. The log-X toggle flips the X
axis to logarithmic, rejects rows with dose ≤ 0, and samples
the fit curve in log space.
Box plot
Distribution per group with Q1 / median / Q3 boxes, 1.5 × IQR whiskers, optional jitter overlay, and optional outlier markers. A per-group descriptive table in the stats panel lists group · n · median · IQR · outlier count. Notched boxes are optional. Non-overlapping notches imply different medians at ~95 % confidence.
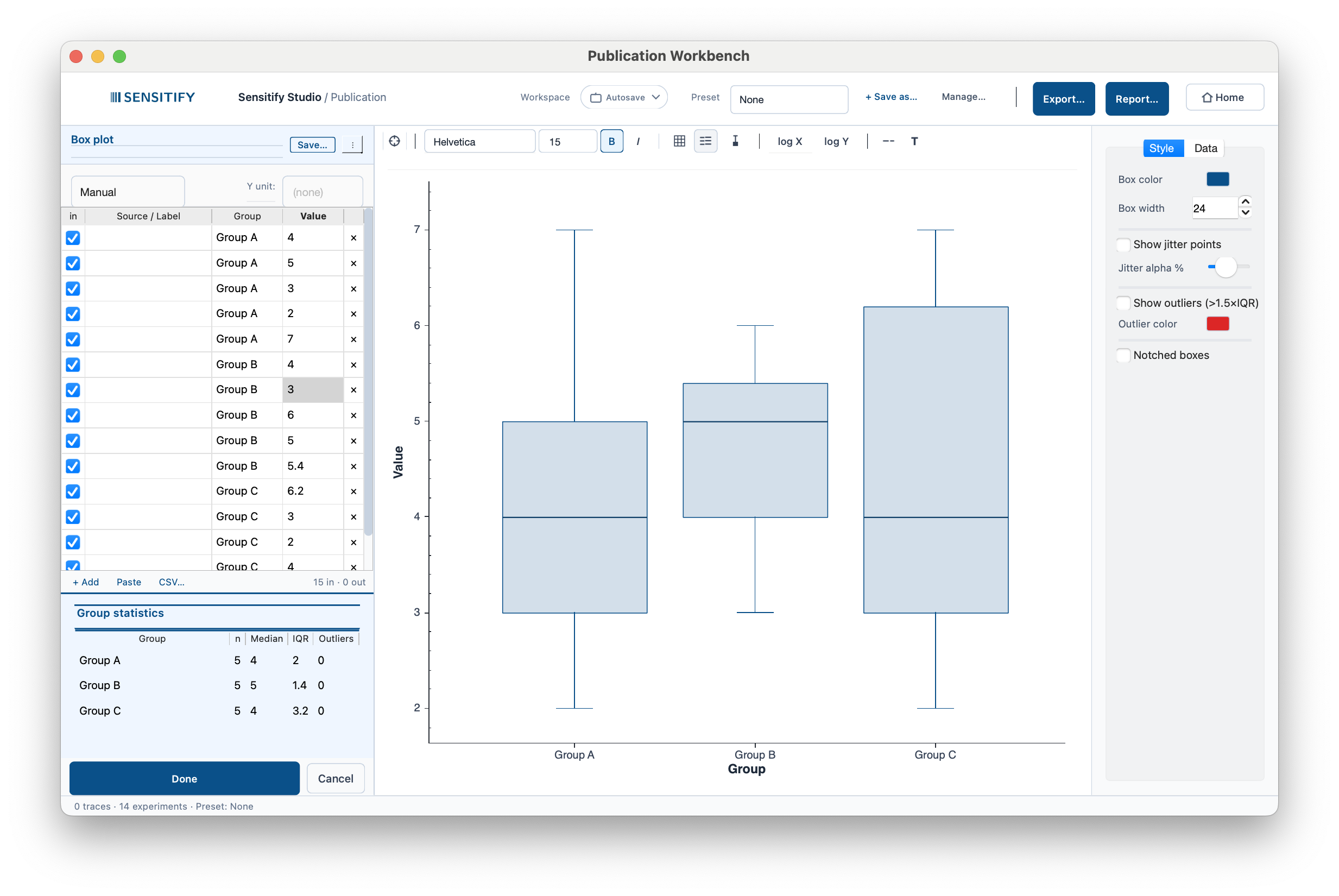
Three groups (n = 5 each), median values around 4–5, with IQRs of 1.4–3.2. The Style panel on the right exposes box width, jitter overlay opacity, the 1.5 × IQR outlier threshold, and the notched-boxes option.
Bar graph
The bar-graph workflow is a 3-tab modal dialog (Data / Style / Statistics) with its own live preview pane and named presets. The modal pattern works here because iteration density is lower than in the in-canvas modes. You set up groups, add significance brackets, and commit.
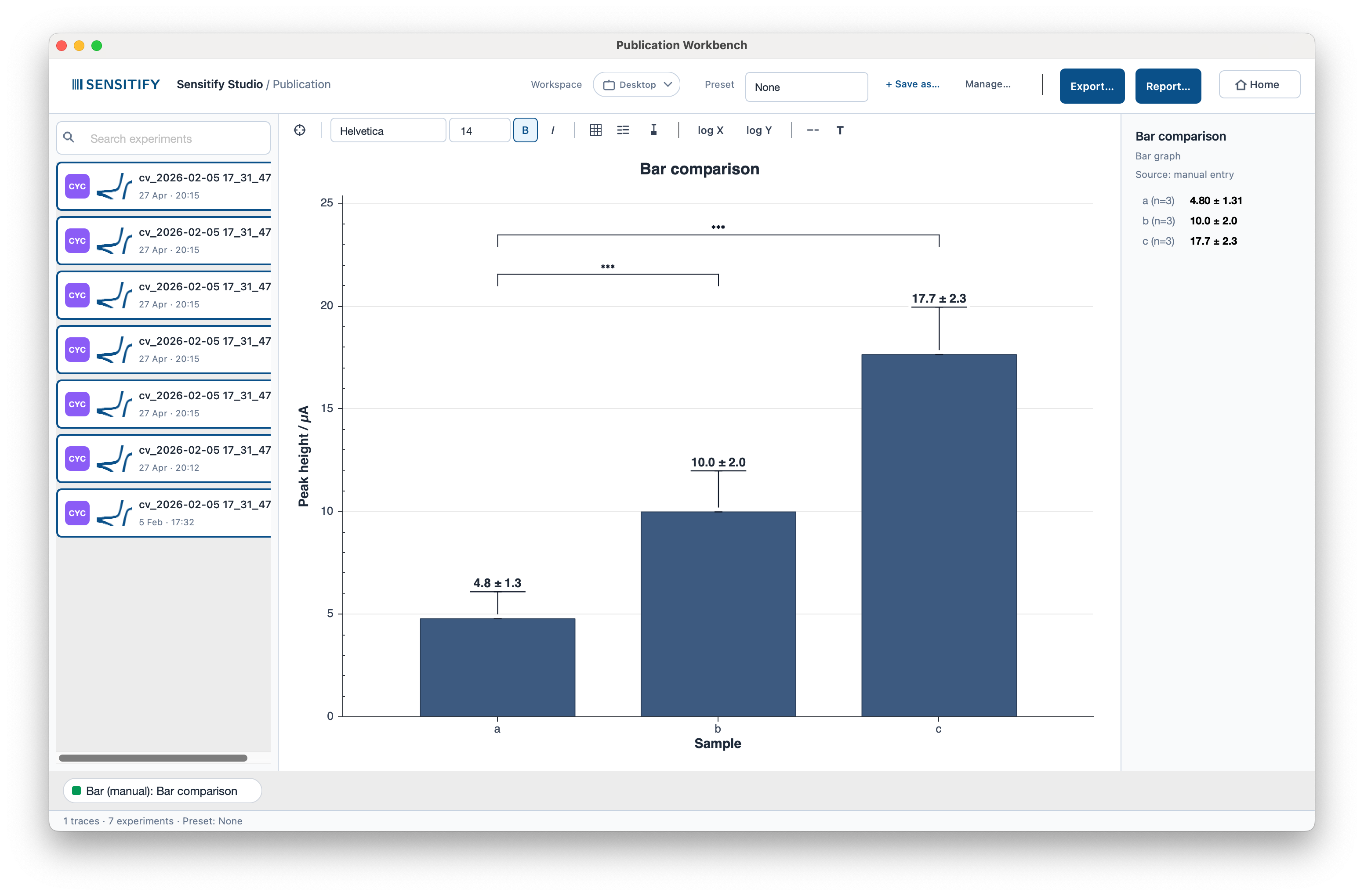
A bar comparison of three samples (n = 3 each) with means, standard deviations, and significance brackets. Values come from the experiments in the left rail or from manual entry, and the dialog keeps two independent buffers so switching modes never loses what you typed. Bar widths adapt to group count (2 → 0.50, 3–5 → 0.60, 6+ → 0.70).
Multi-panel and Blank canvas
Two layout containers round out the welcome grid:
- Multi-panel. Composite layout of several plots in a journal-style grid. Aligns axes, shares legends, and generates panel labels (a / b / c / …).
- Blank canvas. Free-form drag-and-drop figure builder for layouts none of the templates cover.
Publication presets
The preset selector at the top of the window switches between five built-in publication-house styles plus user-saved slots:
| Preset | Family | Title | Axis | Tick | Frame | Notes |
|---|---|---|---|---|---|---|
| None | Helvetica | 18 bold | 15 bold | 13 | 1.2 px | Sensitify house style. |
| Nature | Helvetica | 17 bold | 15 bold | 13 | 1.2 px | No top / right frame. Nature submission palette. |
| IEEE | Times New Roman | 17 bold | 15 bold | 13 | 1.3 px | All-frame box, greyscale palette. |
| ACS | Helvetica | 19 bold | 16 bold | 14 | 1.5 px | Tableau-style palette, ACS column widths. |
| Presentation | Arial | 26 bold | 21 bold | 18 | 2.2 px | Slide-deck sizes, ~50 % bigger. |
Switching preset re-skins every trace on the canvas in one click. It also drives the in-canvas modes' default scatter / fit colours and line widths, so a Nature-styled calibration curve looks like a Nature-styled Tafel plot. Manually overridden per-trace colours are preserved; the preset does not clobber your choices. Save as… captures the current state as a named user slot. Manage… lists saved slots with rename / delete.
Live fit, two-way binding
Every in-canvas mode runs the fit 100 ms after each table edit. The plot and the stats panel both reflect the new values without a refresh button anywhere.
The plot ↔ table link goes both ways. Click a point in the plot and the table scrolls to that row and highlights it. Click a row in the table and the matching point gets a ring on the canvas. To exclude an outlier, right-click the suspect point and choose Exclude from fit. The fit re-runs without it; R² and slope update in the panel within 100 ms.
In Origin, the same outlier round-trip is a workbook scroll, a column edit, a script re-run, and a manual figure refresh. In Excel, it is a trendline that breaks every time you sneeze. The in-canvas pattern keeps the iteration in the visual flow.
Workspaces
The Workspace chip in the header switches between saved project workspaces. Each workspace is a directory containing the experiment library plus the figure session for that project, paper, or experimental campaign. Autosave is the always-available scratchpad workspace.
Export and Report
- Export writes the current figure as PNG, PDF, or SVG. PDF for manuscript figures (vector), SVG for editable hand-off to a designer, PNG for slide decks and posters.
- Report assembles a multi-figure PDF from every figure in the current Workbench session. Use it to share a whole campaign with a collaborator or attach it to a lab notebook.
